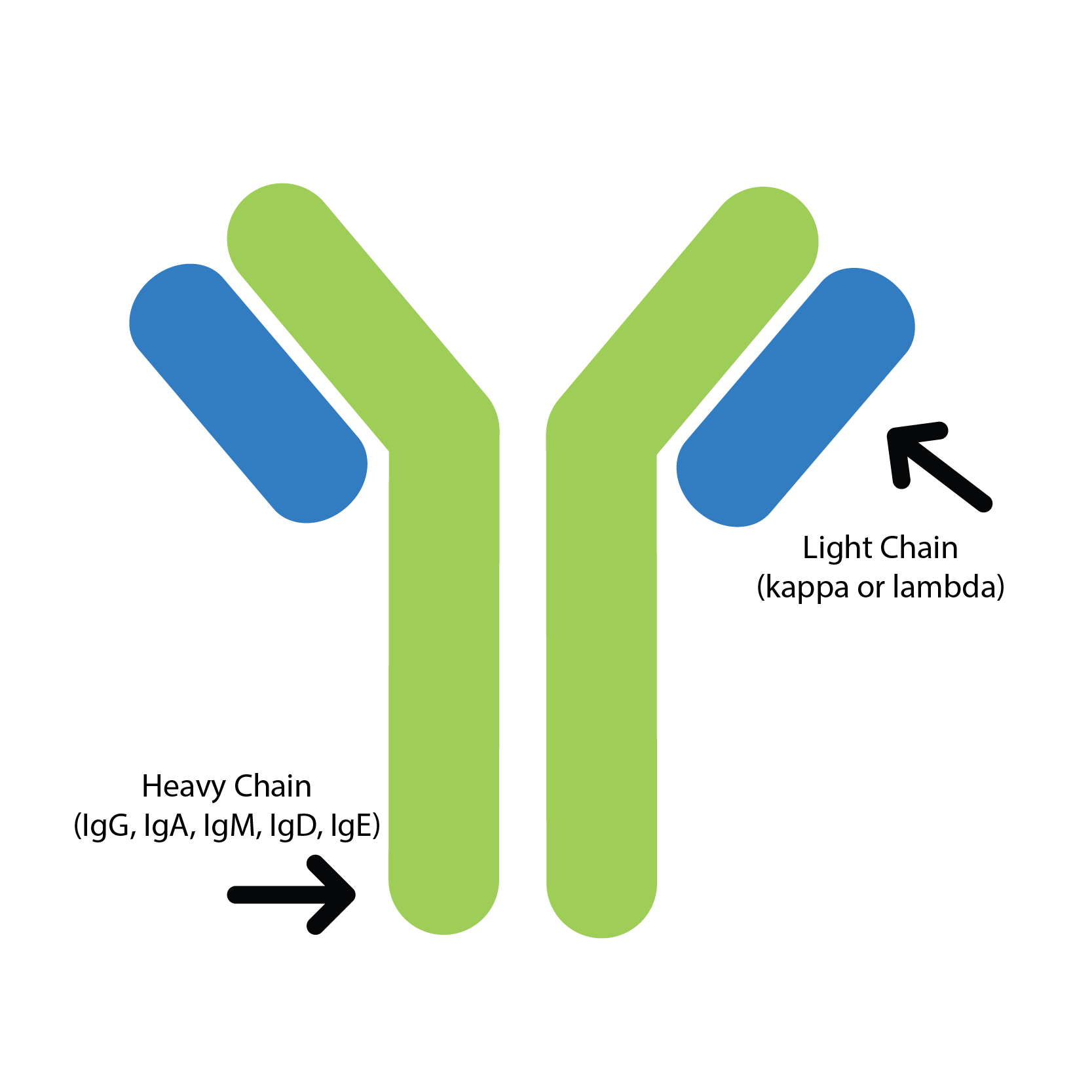
Measurement of kappa (κ) and lambda (λ) light chains in free and total... | Download Scientific Diagram
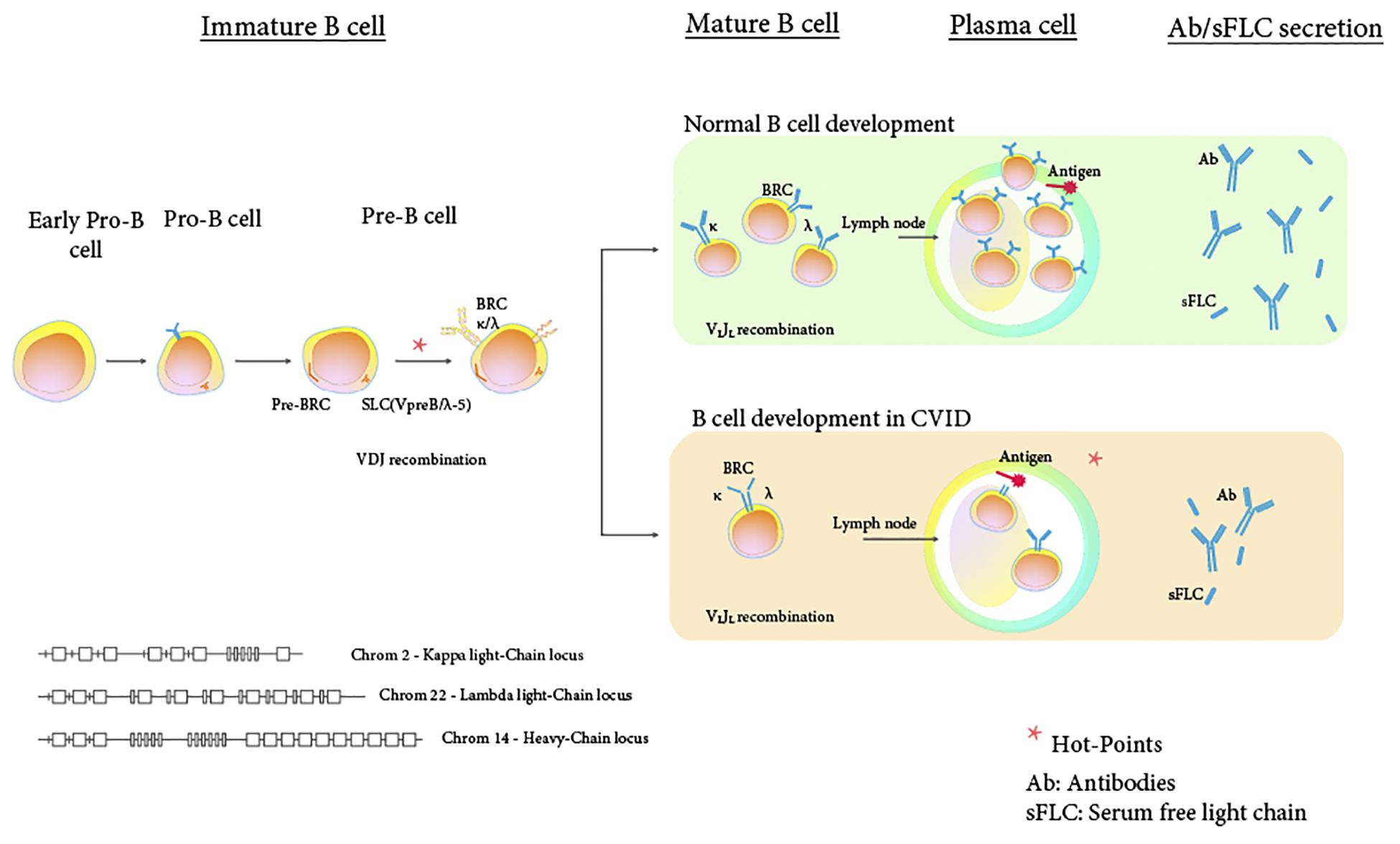
Frontiers | Serum Free Immunoglobulins Light Chains: A Common Feature of Common Variable Immunodeficiency?
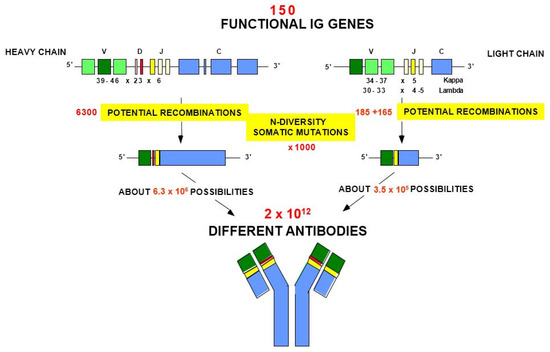
Biomedicines | Free Full-Text | Immunoglobulins or Antibodies: IMGT® Bridging Genes, Structures and Functions
Excluding myeloma diagnosis using revised thresholds for serum free light chain ratios and M-protein levels | Haematologica

Paired heavy- and light-chain signatures contribute to potent SARS-CoV-2 neutralization in public antibody responses - ScienceDirect
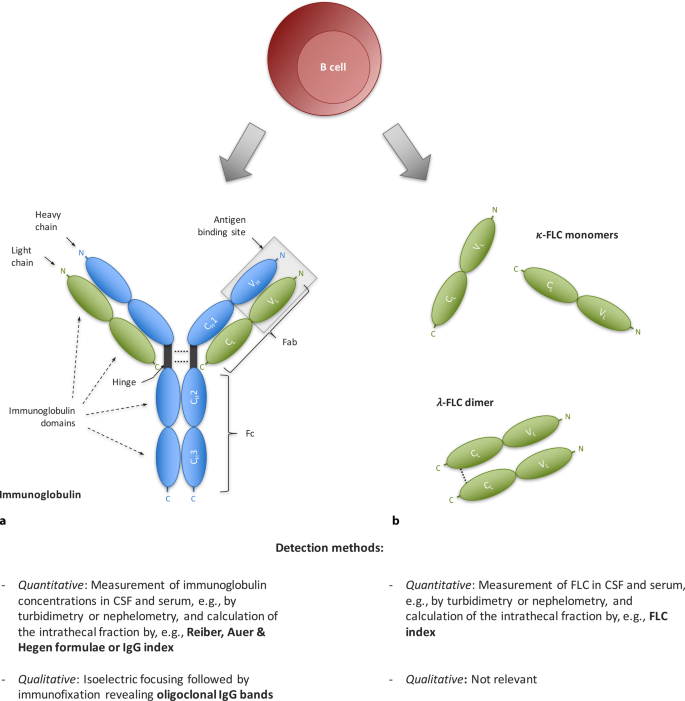
Cerebrospinal fluid kappa free light chains as biomarker in multiple sclerosis—from diagnosis to prediction of disease activity | SpringerLink

What is the Difference Between Kappa and Lambda Light Chains | Compare the Difference Between Similar Terms
Mutations in Specific Structural Regions of Immunoglobulin Light Chains Are Associated with Free Light Chain Levels in Patients with AL Amyloidosis | PLOS ONE
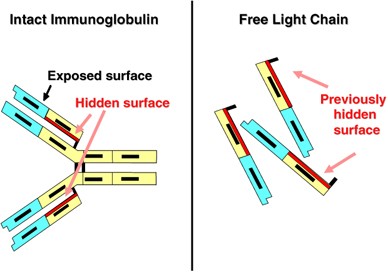
International Myeloma Working Group guidelines for serum-free light chain analysis in multiple myeloma and related disorders | Leukemia

NF-kB Nuclear Factor Kappa-light-chain-enhancer of Activated B Cells Protein Complex. Plays a Role in Cancer and Inflammation Stock Illustration - Illustration of complex, protein: 187955906

Kappa-on-Heavy (KoH) bodies are a distinct class of fully-human antibody-like therapeutic agents with antigen-binding properties | PNAS

Zach London on Twitter: "6/ Different heavy and light chains are associated with different malignancies. Get to know which with this handy chart. As for me, my heart beats a little faster
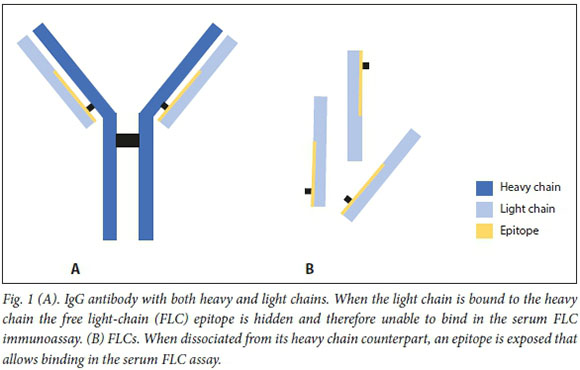
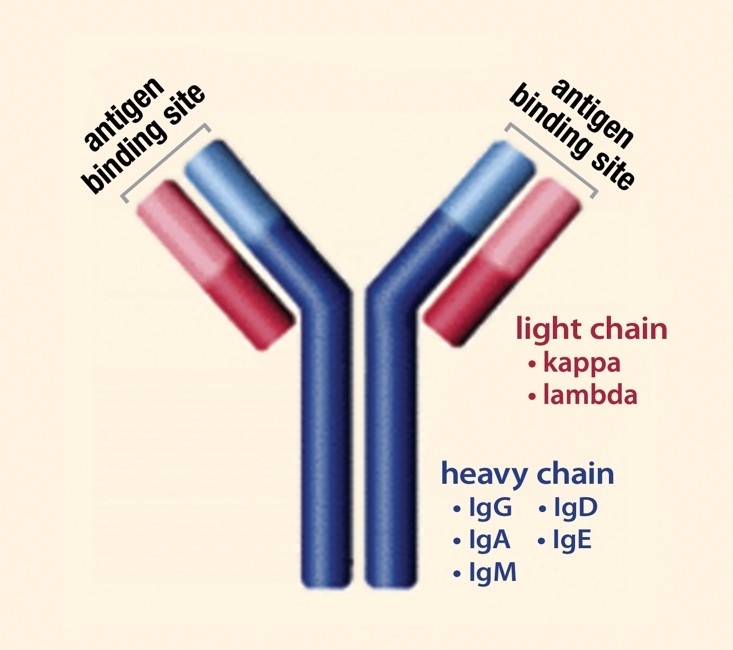

![Native Human KAPPA LIGHT CHAIN Protein [229-20237] Native Human KAPPA LIGHT CHAIN Protein [229-20237]](http://www.raybiotech.com/images/detailed/77/Protein_button.gif)


![Human lambda light chain Antibody (1000011) [Unconjugated] (MAB10049): Novus Biologicals Human lambda light chain Antibody (1000011) [Unconjugated] (MAB10049): Novus Biologicals](https://images.novusbio.com/images2/Ig_Lambda_Light_Chain_MAB10049_Simple_Western_23184.jpg)